Scientists just designed a functioning genome with the minimum number of genes needed for life—and they still don’t understand a third of what those genes are supposed to do.
To define the exact functions of specific genes, experts are looking at some of the simplest creatures alive. Researchers at the J. Craig Venter Institute in San Diego, California are deconstructing the genomes of these modest bundles of life and then stitching them back together in an attempt to find the minimum number of genes necessary for survival.
In 2010, Craig Venter and Clyde Hutchison synthesized the genome of
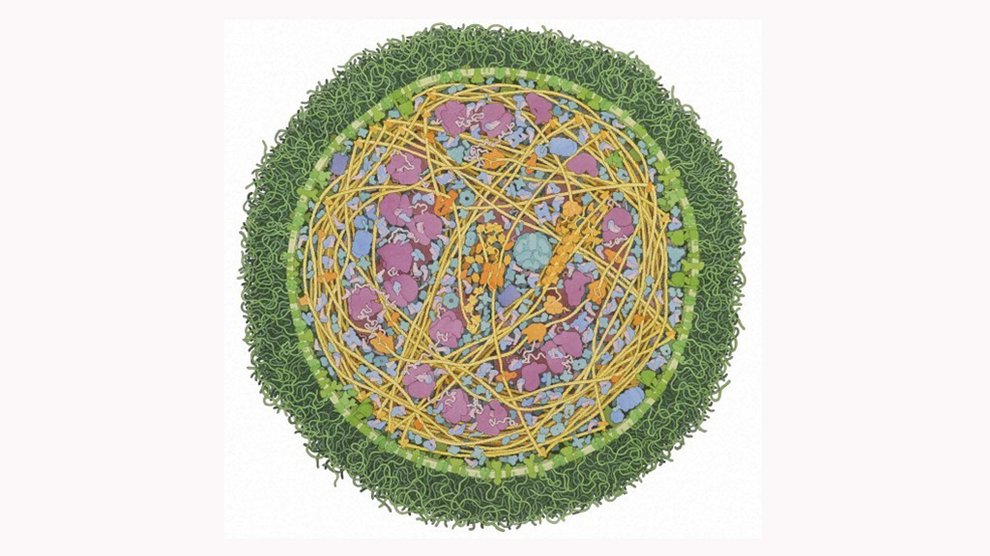
To minimize Syn 1.0, Venter and Hutchinson spent five years inserting transposons—foreign genetic sequences—into various genes to disrupt their operations and see if they’re actually required for survival. They also hunted for quasi-essential genes, which are necessary for the bacterium’s optimization but not for pure existence. After performing these tests on the genome in eight different “chunks,” the researchers came up with Syn 3.0, a minimal viable
M. mycoides genome composed of 473 genes. They present their results today in the journal Science.Syn 3.0 isn’t the definitive minimal viable genome out there—it’s just the most basic we’ve been able to create so far. The team also emphasizes that every genome is context-specific; an organism with a different metabolism, for example, would have a very different minimal genome. But synthesizing a minimal cell is the first step in retracing the billions of years of genetic iterations that went into the emergence of more complex life. “In theory, we could add gene sets and essentially create any organism,” Venter said.
Of the minimum genome’s 473 genes, there are 149 that scientists don’t yet understand the precise function of. “Knowing that we’re missing a third of our fundamental knowledge is a very key finding,” Venter said. It means that experts need to focus on these genes (which encode possibly universal proteins) before they can move onto more complicated collections of genes.
The finding could not only help illuminate what we don’t know about the genome—it could also have industrial applications. For example, if we can build whole genomes using an expanded repertoire of genetic pathways using chemical synthesis, scientists may be able to generate microbes that can pump out medicines, biochemicals, or biofuels.
Venter says the finding has solidified his sense that life is based on the overall workings of the genome, rather than the individual genes an organism is made of.
“Life is much more like a symphony orchestra than a piccolo player,” Venter said. “We’re applying the same philosophy now to our analysis of the human genome. Most human conditions are affected by variations across the entire genome.”
In other words, he says, there’s no “gene for X”; the genome as whole dictates genes’ synchronization—and an organism’s phenotype, as a result. Syn 3.0 may be the first landmark development in a new era of synthetic biology that will further this uncharted territory.


