When I arrived at the remote clinic some 180 miles from Nairobi, it was near mid-day, and the Kenyan sun had pushed the thermometer to uncomfortable heights. Small insects buzzed in and out of the open windows of the low building—the only thing keeping them from landing on the patients were the insect nets draped over each bed. The medical staff at Kimalel Health Center have limited resources available. They don’t have the wide-range of medicines that are available in developed countries, and their equipment is often old and on the edge of disrepair. Despite this, they went about their business with cheerful efficiency.
Gathered in the waiting area was a small collection of families. Young children around the same age as my own kids stared at us as we walked in. Some of these families had walked for days to get to here, a clinician had said. Their kids had symptoms of kala-azar , or visceral leishmaniasis, a disease caused by a parasite that causes fever, swelling of the spleen and liver, and, if untreated, can lead to death. Kimalel is a field site for clinical trials run by the Drugs for Neglected Diseases Initiative, and it was well known in the region as the place to get treatment.
For patients, getting to the clinic is only the beginning of what is potentially a very long journey. The only way these infections can be conclusively diagnosed is by snipping with a needle a small bit of spleen from the patient’s abdomen. The drugs used to treat the confirmed cases are painful and toxic and require hospitalization for a month. Looking at those families in the waiting room, I could see the faces of my own kids. My own worry was reflected on each parent’s face.
Leishmaniasis is what’s known as a neglected disease. Because it primarily infects the poor who cannot pay for expensive treatments, there are few incentives for the pharmaceutical industry to develop new and better drugs. The drug discovery process these days is high-risk and high-cost, as much as $1 billion per new drug. There’s simply no way that a company can recoup the costs by selling drugs that patients can’t pay for.
I have seen the drug discovery process first hand, having worked in the pharmaceutical industry for a number of years. Its mission to treat illnesses energized me, but I soon realized there were a number of diseases that didn’t fit a particular business model. Still, I felt strongly that the way companies discover drugs for cancer or diabetes could be equally fruitful when applied to neglected diseases. So about six years ago I left my job at a company to work at a university. I wanted to do the same sort of work—drug discovery—but in a non-profit environment where I could focus on leishmaniasis and other neglected diseases that are caused by protozoan parasites, such as African sleeping sickness, Chagas disease, and malaria. These insect-borne diseases affect a wide range of populations around the world, and yet there’s a paucity of drugs to treat them.
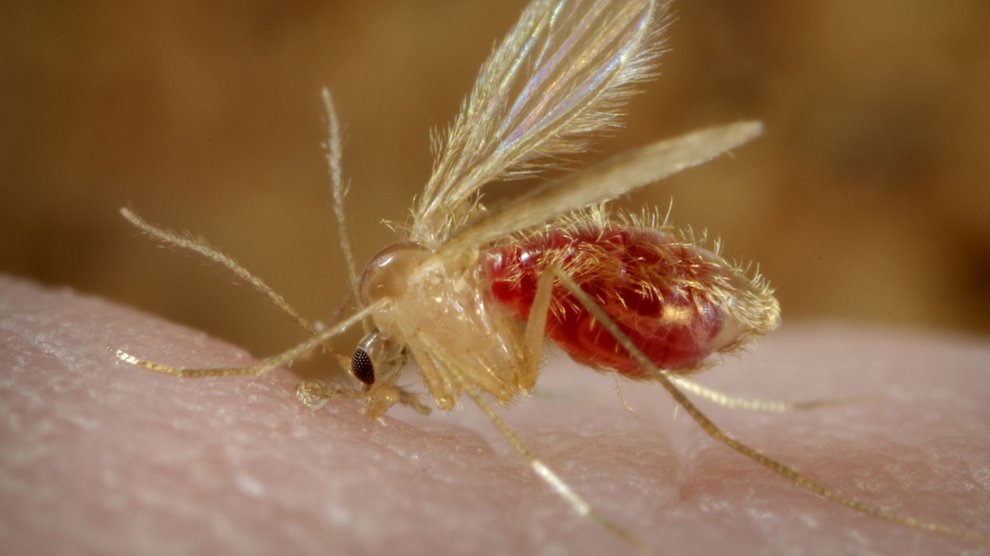
Neglected diseases are anything but rare. Across the globe, about a billion people have one of these neglected diseases, though there are disproportionally few research labs working to discover new drugs for them. Part of the problem is the relative lack of funding: The total, world-wide research and development expenditures for the 31 most common tropical diseases is about $3 billion dollars. Nearly two-thirds of this is spent on the “big three” diseases: HIV/AIDS, malaria, and tuberculosis. That leaves only about a billion dollars for the remaining 28. The economic crisis and government sequester in 2013 has only made the situation worse, restricting the ability of the National Institutes of Health—a primary funder of neglected disease research—to support such work. As a result, there is not much money available to advance discovery of new drugs.
There’s little chance that’s going to change in the near future, either, which simply means my colleagues and I will have to do more with less. We are hoping to overcome this problem in much the same way as the clinicians in Kamalel address their funding limitations: by finding more pragmatic, thrifty, and efficient approaches.
Filling the Gaps
The typical drug discovery process is lengthy and laborious, often beginning by screening hundreds of thousands of compounds at once for candidates with potential. The list is then slowly winnowed down over the course of months or years until the most promising compound is left. Finally, the top choice is optimized for potency and specific properties.
In order to trim this sizable—and expensive—process, my laboratory is taking a more targeted approach. Rather than sift through hundreds of thousands of compounds, we start with existing ones that were developed for other diseases, like cancer. Many of the enzymes and biological pathways responsible for a parasite’s activity are very similar to those found in human cells. That means we can use established drugs that inhibit human enzymes as starting points. With some modifications, they can be used to restrict the growth of parasites that cause neglected diseases. So far, we’ve had some success in our search for new antiparasitic drugs with FDA-approved drugs like lapatinib, a breast cancer drug, or compounds that are still under development.
The process typically goes something like this: First, we test these drugs against lab-grown cultures of the parasite. We study how the drug affects the bug and then design and synthesize new variants based on the results. We work to improve the existing drug’s efficacy, try to reduce its toxicity to humans, and devise a dosing regimen specific to the disease. By the end, the old compound has morphed into a new drug that is uniquely designed for a specific neglected disease.
It’s a much leaner, more streamlined process, but there are still points at which we need assistance. The pharmaceutical industry, despite not spearheading drug discovery for neglected diseases, has built significant research infrastructure, and they’ll occasionally grant access to their facilities for labs like mine. Thus, while the pharmaceutical industry does not directly fund our research, their substantial infrastructure has provided us with significant experimental and consulting support that has been indispensable.
For example, we collaborated with GlaxoSmithKline and their “ OpenLab ” in Spain. The company performed a 45,000-compound high-throughput screen of a subset of their corporate screening set against Trypanosoma brucei , the parasite that causes sleeping sickness. The data they provided to us has spawned several drug discovery projects in our laboratory, and the company continues to collaborate with us. The OpenLab has also pursued a large number of similar projects against other parasitic diseases, such as leishmaniasis, malaria, and tuberculosis, providing significant financial support and offering their expertise. After the tests are finished, they deliver the resulting compound data to academic collaborators like myself to follow up on. Importantly—and perhaps surprisingly—under the OpenLab program, GSK does not demand intellectual property rights for new drugs developed through the program. Rather, it views this contribution of resource and effort to be a matter of corporate responsibility—doing the right thing.
In another example, AstraZeneca has provided us with advice and experimental support to study how drugs are metabolized by the body, which consists of a series of highly specialized and costly tests that are difficult to complete in the academic environment. Such studies are critical to the process of tweaking a compound for safety and efficacy—they help us understand how the body will process a drug. The company has agreed to allocate some of its excess capacity for automated drug metabolism studies to perform experiments in support of our neglected disease work.
Sharing the Results
There’s a caveat to this, though. High-throughput screening and drug metabolism studies produce copious amounts of data. In fact, any one drug-discovery project generates huge volumes of data as new compounds are synthesized, tested for antiparasitic activity and toxicity, and potentially advanced into animal studies. So much, in fact, that it demands a new model to more effectively share data between different teams.
Within a single drug company, this data would be archived and available to other project teams. But in the academic environment, labs around the world don’t yet have many good ways to quickly share data and learn from each other’s experiments. Typically, experimental results are still shared the old fashioned way—in research publications and presentations at scientific conferences. The delay between an experimental result and the release of its data can be many years or months, making the traditional publication process something of an anachronism given the fast pace of today’s drug discovery process.
Because of this, it’s very likely that neglected disease drug discovery labs around the world are unknowingly performing the exact same experiments simultaneously. With such limited resources available to neglected disease research, this duplication of effort is unacceptable. It’s something that I and others have been working to remedy. For example, my lab recently secured funding from a crowdfunding campaign to build a data sharing system for neglected disease drug discovery. Using this system, labs from around the world will share their experimental data with each other. We believe strongly that this will reduce duplication of effort and allow ideas to flow more freely between labs.
The challenges, though, are formidable. Such data sharing would be a significant culture shift among drug discovery scientists, who have often been trained to keep data secret to ensure competitiveness and the ability to file patents (which can translate to financial benefits). However, while this may be true for certain diseases that primarily affect patients in the developed (read: rich) parts of the world, I don’t believe that such a model is relevant or even appropriate for these neglected diseases, which disproportionally affect the poor.
Over the last several years, I have grown more and more certain of the critical role that the academic and non-profit world must play in discovering new drugs for neglected diseases. Drug discovery was once mostly relegated to the pharmaceutical industry, where vast expertise and experimental resources are available for the highly complex process. Now, those same capabilities are increasingly available in the university environment. Neglected diseases are an area where we can make a tangible difference.
However, we cannot simply replicate the way that pharmaceutical companies discover drugs—we simply don’t have the funds. Rather, we must take a cue from the clinicians in Kimalel, finding creative and cost-effective ways to perform our work. By sharing data and applying new approaches to drug discovery, we can accelerate the path to new drugs for the neglected patients who so urgently need them.


