For more than two years, Ebola stalked West Africa, sickening nearly 30,000 people and killing almost 12,00 more. In each person Ebola infected, the virus copied its genetic material billions of times by sneaking into countless cells in every major organ system. As cells filled with copies of the virus, they burst open, spilling their contents and letting the cycle begin again. For those infected, the result was hemorrhage and, frequently, death.
That it kills such a large proportion of people it infects in chillingly gruesome ways gives the virus an air of the macabre. Even after nearly four decades of study, the Ebola virus remains shrouded in mystery. Scientists still don’t know the animal species in which Ebola lurks between outbreaks. Nor do they know how or why the virus first popped up in Guinea in December 2013, more than 1,500 miles from where it was first discovered in Uganda nearly 40 years before.
Chris Hempel hadn’t given much thought to Ebola, as her attention was consumed by her job as full-time caregiver for her 11-year-old twin daughters Addi and Cassi, who have a rare disease called Niemann-Pick Type C (NPC). The disease, sometimes referred to as “childhood Alzheimer’s,” interferes with the cell’s ability to process cholesterol and causes brain damage and early death. With only 500 people affected by NPC worldwide, studies to understand and treat the disease were almost as rare as the disease itself. Sitting at her laptop at her Reno, Nevada, home in 2011, however, Hempel received a stack of news articles in her inbox about Ebola that appeared alongside the usual load of Niemann-Pick news. As she read deeper, she realized that Kartik Chandran, a virologist at Albert Einstein College of Medicine in New York, and colleagues had discovered that, to enter cells, the Ebola virus used the same protein that was mutated in Addi and Cassi, NPC1.
For virologists, it was a key discovery in understanding the molecular biology of Ebola. For Chandran, Hempel, and other NPC parents, it marked the start of a collaboration that would help crack open some of the deepest mysteries of one of the world’s deadliest viruses.
Lethal Origins
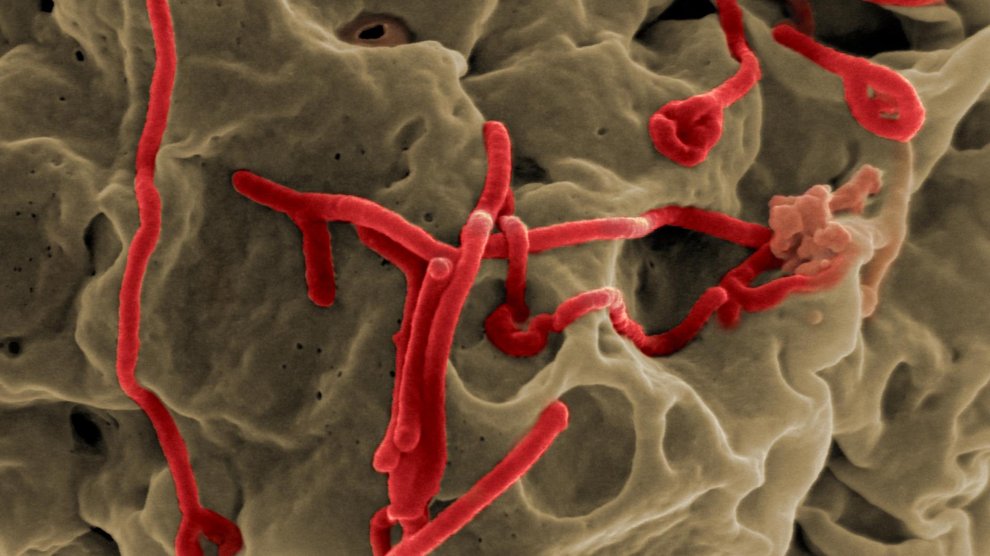
Ebola first emerged in 1976 in the tiny village of Yambuku along the river after which the virus was named. When virologists first peered through their microscopes to see the cause of the illness that killed upwards of 90% of the people it infected, they captured the iconic image of the filovirus that now fills popular and scientific literature. The Ebola virus itself doesn’t look inherently sinister. Unlike other viruses that take on complex geometric shapes, Ebola looks like a tiny piece of spaghetti carelessly left on the plate.
Its genome was small, only 18,960 RNA nucleotides coding for seven proteins, yet the virus could infect nearly every type of cell in the body. Even as scientists chipped away at other aspects of Ebola biology—sequencing its genome, understanding how the virus was spread, and identifying antibodies produced in response to infection—they still couldn’t figure out how the virus actually got inside the cell. All animal cells are studded with proteins and sugars that to viruses act as cellular locks. If the virus can evolve the right set of lock picks, it can use a specific cell membrane protein to sneak inside, hijack the cell’s replication machinery, and copy itself.
“Viruses co-opt our own cellular machinery to infect us, which makes those proteins good targets for therapeutics,” Chandran says.
The cellular lock and viral lock pick is so important to understanding the basics of how a virus works that it’s often one of the first things virologists search out. They quickly identified the protein that Ebola used as a lock pick, a viral spike glycoprotein, just as many filoviruses do. But what researchers couldn’t find was the cellular lock that the virus used.
Over the years, virologists sifted through proteins to identify the one that held the key to Ebola. Instead of coming up empty-handed, however, researchers like Chandran had the opposite problem. Their experiments yielded a long list of proteins that the Ebola virus could use. When scientists tried mutating some of these proteins in dishes of human cells, they expected it would change the cellular lock to one the Ebola virus couldn’t pick. It didn’t—the virus broke into the cell just as easily as it had before.
“We were really just stumbling around in the dark for a long time,” Chandran says.
Chandran decided to temporarily ignore all previous work, starting fresh and making no assumptions. His team began with a set of cultured human cells in which they randomly generated mutations, one mutation per set of cells. Rather than working with the entire Ebola virus, which would require painstaking work in a Biosafety Level 4 (BSL4) lab by scientists in full protective gear, they tacked the Ebola viral spike glycoprotein onto a much more benign virus, the vesicular stomatitis virus that infects livestock. The engineered virus would, in theory, be able to unlock all of these cells except for the one in which the lock was broken.
Their first pass yielded a result no one expected. The only cellular mutation that affected the virus’s ability to infect the cultured cells was in a protein called NPC1, which helped transport cholesterol within the cell. NPC1 was not expressed on the surface of the cell—where you might expect a lock to be—but rather on small inner compartments called lysosomes and endosomes. It was like finding out that a burglar had broken in not by unlocking the front door, but one to the bathroom. Yet when Chandran repeated his experiments, he got the same results. Cells with mutations in NPC1 were resistant to infection with the Ebola virus.
“I don’t think any of us, if we were honest, knew jack-all about Niemann-Pick C1,” says John Dye, a virologist at the U.S. Army Medical Research Institute of Infectious Diseases in Fort Detrick, Maryland, who was collaborating on Chandran’s Ebola research. “In a million years, I never would have picked this protein.”
Chandran he turned to the medical literature to learn more. No one had yet figured out the exact function of the protein, but it appeared to be involved in the movement of cholesterol and other lipids both within and between cells. Without a functioning NPC1 protein, lipids and cholesterol build up inside cells, unable to get where they’re needed in cellular membranes and throughout the body. Too many fats in the wrong places and not enough in the right ones leads to cellular dysfunction and death.
Affected children like Addi and Cassi have cognitive and developmental problems along with difficulties moving and eating. At age 11, they are some of the oldest people in the world with NPC disease.
Chandran didn’t know the Hempels, but he did know the twins’ cells. Not long after Addi and Cassi were diagnosed, Chris, her husband, the twins, and all surviving grandparents donated fibroblast cells to a biobank run by the Coriell Institute in New Jersey. Chandran ordered their cells and those from other NPC families to test the role the NPC1 protein played in Ebola transmission. He passed these cells to Dye, who could test the cells with the real Ebola virus, performing the experiments in the facility’s BSL4 lab. Cells with two defective copies of NPC1, like those obtained from Addi and Cassi, were completely resistant to the Ebola virus. Carriers of the disease—people like Hempel who had one normal copy of NPC1 and one mutated copy—were partially immune.
But that still didn’t explain how the Ebola virus could enter the cell to access the NPC1 protein in the first place. Rather, the answer would lie in how the Ebola virus shielded itself from the immune system.
Typically, when a virus infects a person, their body’s immune system generates antibodies that can gum up the virus particles’ lock picks to prevent them from getting inside cells. Chandran’s previous work showed that the sugar molecules that are attached to the spike proteins on the outside of the virus shielded the proteins from the immune system. To infect a cell, the virus used an enzyme called a cathepsin to remove the sugar molecule. But again, Chandran was left with the same, vexing mystery: how could a virus infect a cell if cathepsin was already in the cell?
Yet once he uncovered the secret of the NPC1 protein, Chandran pieced together a hypothesis of how the Ebola virus might infect human cells. The viral spike glycoprotein, shielded by the sugar molecule, acted like a skeleton key and tricked the cell into swallowing the virus whole in a process called pinocytosis. A cell then shuttled the virus to the lysosome, which is part of the cell’s recycling facility and contains many enzymes, including cathepsin. There, cathepsin neatly snipped the sugar molecule off the viral spike glycoprotein. The naked protein on the surface of the Ebola virus could then bind to the NPC1 protein to exit the lysosome and begin copying its own genetic material. Chandran’s group showed that interfering with this process by blocking either the activity of cathepsin or the binding site of NPC1 prevented Ebola from making copies of itself and spreading between cells.
The team published these results simultaneously in two papers in Nature in 2011, showing the world they had finally found the receptor for the Ebola virus. Journal editors noted that the “unexpected role for the hereditary disease gene NPC1 in Ebola virus infection may facilitate the development of antifilovirus therapeutics.”
The push to create these medicines took on added urgency when the most recent and largest Ebola outbreak began in West Africa in 2013. In their efforts to develop vaccines and antiviral medication, researchers would increasingly turn to children suffering from one of the world’s rarest illnesses.
A Key Protein
When Gayla Chan was born in October 2012, it looked like the newborn herself was pregnant. Her liver and spleen were nearly four times their normal size, pushing out her tiny abdomen and compressing her lungs. Difficulties breathing meant that the infant spent most of her first year of life in the hospital. A genetic test revealed Niemann-Pick Type C. As a computer programmer and software developer, her father Brian Chan immediately turned to the Internet to search for more information on Gayla’s illness and to promote the need for better research on NPC. Chan convinced the entire family to donate cells to the Coriell Institute to further research into finding treatments for NPC.
One such treatment is a molecule called cyclodextrin, a ring-shaped sugar molecule that scientists thought would help remove excess cholesterol from cells. In April 2009, the Hempels filed a compassionate use investigational new drug application with the US Food and Drug Administration to administer cyclodextrin directly to Addi and Cassi’s spinal cords. Hempel’s enthusiasm for the drug spurred researchers to develop new animal models to test the therapy. Early successes lead to the creation of a phase one trial led by the National Institutes of Health. As part of this trial, Gayla Chan began receiving cyclodextrin in June 2014, the youngest child ever treated with this condition.
Chan believed the cells he had donated to Coriell would ultimately be used in researching cyclodextrin or other NPC-related studies. Instead, the first scientists to use his cells were virologists in New York City who were working on Ebola. As Chan read the 2011 Nature studies, he felt one of the first glimmers of hope since his daughter was born. The research might not save Gayla, but it just might save other children. Chan wanted to contribute to Ebola research, so he sent Chandran nine samples of fibroblast cells from skin biopsies taken from himself and his wife, their three children, and both sets of grandparents. Experiments from Chan’s family and others with NPC showed that Ebola grew much slower in people who had Niemann-Pick or were carriers of the condition.
“If I got Ebola, I would be better off than 99% of the population,” Chan says.
Steven Walkley, a specialist on lysosomal diseases such as NPC at Albert Einstein College of Medicine, had spent his career studying the NPC1 protein. He developed cell cultures and animal models of NPC disease, never imagining that one day an Ebola scientist would come knocking. It was his collaboration with Chandran and other Ebola scientists that brought the most attention to his work. “All of a sudden, this protein I had been working on for decades was now front and center in the fight against a famous, terrifying infection,” Walkley says.
The existing banks of cells and animal models meant that Chandran, Walkley, and Dye could dive right in to understanding how Ebola and NPC1 interacted in a living organism. As with the cell cultures, Dye found that mice lacking a functional NPC1 protein were completely immune from Ebola. These mice were not healthy—they had jaundice and neurological problems that mimicked those of Niemann-Pick kids—but no matter how much virus Dye injected, it didn’t replicate and the mice didn’t develop Ebola. That wasn’t the case with mice that had two functional NPC1 genes. They invariably developed the classic Ebola fever followed by hemorrhaging, where blood leaked into their body cavity and out through their eyes and noses. Soon after, they all died.
Heterozygous mice, which carried one mutant copy of NPC1 and one normal copy, were somewhere in between. They tended to get sick with Ebola, but they also tended to survive the infection.
The discovery that Ebola used NPC1 as its receptor solved a huge mystery for researchers. To Dye, it also helped explain why Ebola was so deadly. Different receptors stud different types of cells—it’s part of what makes a liver cell different from a brain cell. It’s also why cold viruses only infect the cells of the upper respiratory tract and norovirus only infects the gastrointestinal tract. Ebola’s initial entry point is non-specific, and all cells also have NPC1 on lysosomes, and all cells have lysosomes. This means the virus can infect any cell type it comes across.
“It’s one of the most beautiful pieces of biology I had seen in a long time,” Walkley says.
This also explains why Ebola is so exquisitely deadly. The virus can make so many copies of itself so rapidly that the immune system finds it nearly impossible to reign in the virus before a person dies.
Work by Dye and Walkley raised an intriguing possibility about the evolutionary origins of NPC disease. Its genesis may be similar to sickle-cell anemia, which is caused by a mutation in the gene that codes for the hemoglobin protein that carries oxygen in our blood. The mutation that causes sickle-cell anemia provides resistance to malaria, both in individuals with the disease and in those who carry only one copy of the mutation. Other rare genetic conditions, such as cystic fibrosis and Tay-Sachs disease, are thought to have similar patterns. Evolutionary biologists believe that these benefits help to maintain the mutations in humans, even though they can be deadly in some cases. NPC may follow a similar pattern.
“It’s certainly a very tantalizing hypothesis,” Chandran says.
What scientists need to determine, however, is how long filoviruses like Ebola have infected humans, and whether these infections occurred frequently enough to create the kind of evolutionary pressures needed to maintain such a deadly mutation. Most Ebola outbreaks had been relatively small and in remote locations, with little information on survivors. Scientists didn’t know how many affected individuals had an NPC1 mutation or whether that mutation affected the chances that they would survive an infection. The most recent outbreak, however, is giving Chandran and his colleagues a chance to test this hypothesis by checking the DNA of survivors to see if there are disproportionate numbers of NPC disease carriers. It’s still too soon to say whether this idea will hold up to scientific scrutiny, but Chandran, Dye, and their colleagues are eagerly awaiting results.
Buying Time with Antibodies
The scientists, however, didn’t merely want to better understand how Ebola caused infections. For them, it was only the first step toward a treatment or vaccine for Ebola, to prevent the massive numbers of infections and deaths like those seen in the West Africa outbreak.
With potential vaccines years away and a need to help individuals already sick with Ebola, researchers have begun to try to create artificial antibodies to help boost the body’s own response to the virus. The most promising immunotherapy is ZMapp, which contains three different antibodies specific to one type of Ebola virus. (Scientists have currently identified five types of Ebola). If an outbreak from four of the other Ebola strains, however, ZMapp would be useless. But all Ebola virus strains, and even closely related filoviruses like the one that causes Marburg fever, use NPC1, which would allow pharmaceutical companies to stock Ebola treatments ahead of any potential outbreak.
Chandran’s breakthrough was realizing that treating Ebola was as simple as inducing a temporary case of Niemann-Pick disease and interfering with the function of the NPC1 protein.
“If we can induce—temporarily—Niemann-Pick disease, it might buy time for the immune system to fight back. If we can keep people from crashing, they might have a shot,” Chandran says.
Their goal was to do that using antibodies. Creating an antibody that would attach itself to a receptor on the outside of the cell was straightforward. The NPC1 protein, however, was inside the cell, which made their task much harder. What Chandran wanted to do was build an antibody that would bind to the Ebola virus, be escorted into the lysosome, and also bind to NPC1. Most single antibodies have a Y shape, with both branches of the “Y” recognizing the same piece of protein, making them what scientists call “monospecific.” In the lab, however, scientists could build a Y-shaped antibody with each arm binding to a different snippet of protein: one on the Ebola virus and one on NPC1. The scientists teamed up with M. Javad Aman at Integrated Biotherapeutics in Rockville, Maryland to build this so-called “bispecific” antibody. Aman’s task was to combine to existing monospecific antibodies into an immunological Trojan horse.
Aman combined an antibody known as FVM09, which binds to the sugar-studded proteins that surround the Ebola virus before it enters the cell, with one known as MR72. Researchers had isolated that antibody, which binds to NPC1, from a man who had survived infection with the Marburg virus. Initial tests with the same engineered, benign vesicular stomachitis virus they had used in previous studies showed it was effective, so Dye tested the bispecific antibody in the maximum containment lab at Fort Detrick with the real thing. His experiments showed that the antibody successfully treated mice infected with one of several strains of Ebola, as well as those infected with Marburg. Their work appeared this past September in Science .
The next step is to test these Trojan horse antibodies in non-human primates and, eventually, humans. To Dye, it’s obvious that without the underlying research on Niemann-Pick disease, their work on Ebola would have been several years behind.
“Those models shaved several years off the process,” Dye says.
Spurred by interest in Ebola, research on Niemann-Pick disease has picked up in recent years as well, taking the project full circle. Addi, Cassi, and Gayla remain waiting for treatments and cures for their extremely rare disease, but Chan and Hempel find it gratifying to be able to help in developing treatments for another deadly condition.
“So many people ask me, ‘Why should I care about kids with such a rare disease?’ ” Hempel says. “And this case has the answer: because proteins affect so many things. If Addi and Cassi could understand this research, I think they would be happy that they helped.”



